| General Information | |
|---|---|
| ZINC ID | ZINC000049035553 |
| Molecular Weight (Da) | 506 |
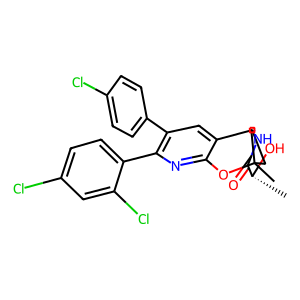
| SMILES | C[C@@H](O)C(=O)N[C@H]1CC(C)(C)Oc2nc(-c3ccc(Cl)cc3Cl)c(-c3ccc(Cl)cc3)cc21 |
| Molecular Formula | C25Cl3N2O3 |
| Action | Inverse Agonist |
| Physicochemical Details | |
|---|---|
| Molar Refractivity | 130.728 |
| HBA | 4 |
| HBD | 2 |
| Rotatable Bonds | 4 |
| Heavy Atoms | 33 |
| LogP | 6.095 |
| Activity (Ki) in nM | 3467.369 |
| Polar Surface Area (PSA) | 71.45 |

| Pharmacokinetic Properties (AdmetSAR) | |
|---|---|
| Human intestinal absorption | + |
| Caco-2 | - |
| Blood brain barrier | + |
| P-glycoprotein inhibitior | + |
| P-glycoprotein substrate | - |
| Cyp3a4 substrate | + |
| Cyp2c9 substrate | - |
| Cyp2d6 substrate | - |
| Cyp3a4 inhibition | - |
| Cyp2c9 inhibition | + |
| Cyp2c19 inhibition | + |
| Cyp2d6 inhibition | - |
| Cyp1a2 inhibition | - |
| Acute oral toxicity | + |
| Carcinogenicity (binary) | - |
| Ames mutagenesis | - |
| Human ether-a-go-go-related gene inhibition | + |
| Biodegradation | - |
| Glucocorticoid receptor binding | + |
| Thyroid receptor binding | + |
| Androgen receptor binding | + |
| Plasma protein binding | 0.786 |

| Pharmacokinetic Properties (SwissADME) | |
|---|---|
| Number of aromatic heavy atoms | 18 |
| Fraction csp3 | 0.28 |
| Ilogp | 3.57 |
| Xlogp3 | 5.97 |
| Wlogp | 6.15 |
| Mlogp | 4.16 |
| Silicos-it log p | 6.6 |
| Consensus log p | 5.29 |
| Esol log s | -6.81 |
| Esol solubility (mg/ml) | 0.0000782 |
| Esol solubility (mol/l) | 0.00000015 |
| Esol class | Poorly sol |
| Ali log s | -7.25 |
| Ali solubility (mg/ml) | 0.0000287 |
| Ali solubility (mol/l) | 5.67E-08 |
| Ali class | Poorly sol |
| Silicos-it logsw | -9.69 |
| Silicos-it solubility (mg/ml) | 0.0000001 |
| Silicos-it solubility (mol/l) | 2.06E-10 |
| Silicos-it class | Poorly soluble |
| Pgp substrate | |
| Log kp (cm/s) | -5.15 |
| Lipinski number of violations | 2 |
| Ghose number of violations | 3 |
| Veber number of violations | 0 |
| Egan number of violations | 1 |
| Muegge number of violations | 1 |
| Bioavailability score | 0.17 |
| Pains number of alerts | 0 |
| Brenk number of alerts | 0 |
| Leadlikeness number of violations | 2 |
| Synthetic accessibility | 4.32 |

| Pharmacokinetic Properties (ADMETLab) | |
|---|---|
| Logs | -5.536 |
| Logd | 4.447 |
| Logp | 6.299 |
| F (20%) | 0.004 |
| F (30%) | 0.005 |
| Mdck | 8.77E-06 |
| Ppb | 1.0071 |
| Vdss | 0.679 |
| Fu | 0.0112 |
| Cyp1a2-inh | 0.569 |
| Cyp1a2-sub | 0.892 |
| Cyp2c19-inh | 0.723 |
| Cyp2c19-sub | 0.079 |
| Cl | 4.723 |
| T12 | 0.034 |
| H-ht | 0.97 |
| Dili | 0.936 |
| Roa | 0.085 |
| Fdamdd | 0.985 |
| Skinsen | 0.031 |
| Ec | 0.003 |
| Ei | 0.006 |
| Respiratory | 0.317 |
| Bcf | 3.761 |
| Igc50 | 5.211 |
| Lc50 | 7.62 |
| Lc50dm | 6.895 |
| Nr-ar | 0.039 |
| Nr-ar-lbd | 0.191 |
| Nr-ahr | 0.801 |
| Nr-aromatase | 0.856 |
| Nr-er | 0.683 |
| Nr-er-lbd | 0.191 |
| Nr-ppar-gamma | 0.509 |
| Sr-are | 0.798 |
| Sr-atad5 | 0.158 |
| Sr-hse | 0.545 |
| Sr-mmp | 0.903 |
| Sr-p53 | 0.94 |
| Vol | 474.363 |
| Dense | 1.063 |
| Flex | 0.208 |
| Nstereo | 2 |
| Nongenotoxic carcinogenicity | 1 |
| Ld50 oral | 0 |
| Genotoxic carcinogenicity mutagenicity | 0 |
| Surechembl | 0 |
| Nonbiodegradable | 3 |
| Skin sensitization | 1 |
| Acute aquatic toxicity | 1 |
| Toxicophores | 2 |
| Qed | 0.389 |
| Synth | 3.665 |
| Fsp3 | 0.28 |
| Mce-18 | 86.062 |
| Natural product-likeness | -0.015 |
| Alarm nmr | 0 |
| Bms | 1 |
| Chelating | 0 |
| Pfizer | 1 |
| Gsk | Rejected |
| Goldentriangle | Rejected |